The paper “Biomedical Relation Extraction with Knowledge Graph-based Recommendations”, authored by LASIGE’s Ph.D. student Diana Sousa has been published in the IEEE Journal of Biomedical and Health Informatics, a top-ranked journal (h5-index 125; Scimago Q1). The paper’s co-author is LASIGE’s integrated researcher Francisco M. Couto.
Biomedical Relation Extraction (RE) systems identify and classify relations between biomedical entities to enhance our knowledge of biological and medical processes. Most state-of-the-art systems use deep learning approaches, mainly to target relations between entities of the same type, such as proteins or pharmacological substances. However, these systems are mostly restricted to what they directly identify on the text and ignore specialized domain knowledge bases, such as ontologies, that formalize and integrate biomedical information typically structured as direct acyclic graphs. On the other hand, Knowledge Graph (KG)-based recommendation systems already showed the importance of integrating KGs to add additional features to items. Typical systems have users as people and items that can range from movies to books, which people saw or read and classified according to their satisfaction rate.
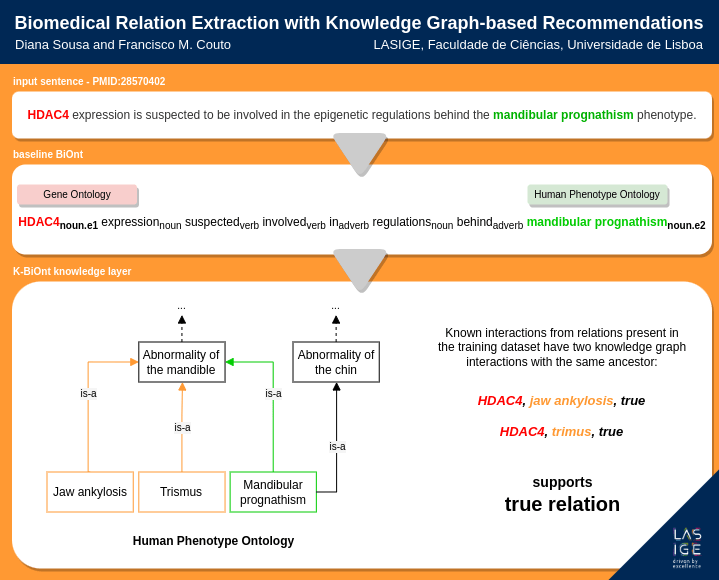
This work proposes to integrate KGs into biomedical RE through a recommendation model to further improve their range of action. We developed a new RE system, named K-BiOnt, by integrating a baseline state-of-the-art deep biomedical RE system with an existing KG-based recommendation state-of-the-art system. Our results show that adding recommendations from KG-based recommendation improves the system’s ability to identify true relations that the baseline deep RE model could not extract from the text. The code supporting this system is available.
The paper is available in early access here.