The paper “G-Tric: generating three-way synthetic datasets with triclustering solutions”, authored by LASIGE’s MSc student João Lobo was published in BMC Bioinformatics, a top-ranked journal. The paper co-authors are LASIGE’s integrated researcher Sara C. Madeira and Rui Henriques from INESC-ID.
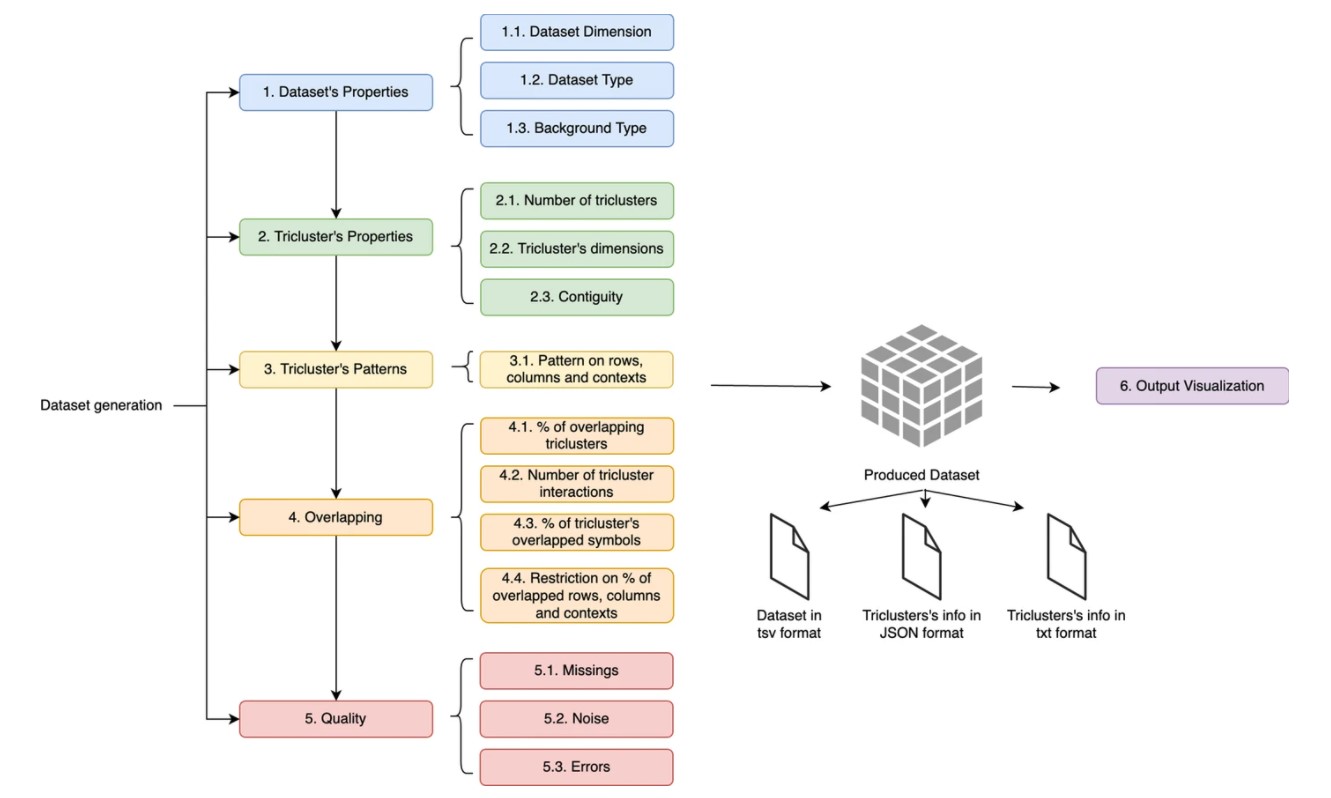
Three-way data started to gain popularity due to their increasing capacity to describe inherently multivariate and temporal events, such as biological responses, social interactions along time, urban dynamics, or complex geophysical phenomena. Triclustering, subspace clustering of three-way data, enables the discovery of patterns corresponding to data subspaces (triclusters) with values correlated across the three dimensions (observations x features x contexts). With increasing number of algorithms being proposed, effectively comparing them with state-of-the-art algorithms is paramount. In this context, G-Tric can replicate real-world datasets and create new ones that match researchers needs across several properties, including data type (numeric or symbolic), dimensions, and background distribution. Users can tune the patterns and structure that characterize the planted triclusters (subspaces) and how they interact (overlapping). Data quality can also be controlled, by defining the amount of missing, noise or errors. Triclustering evaluation using G-Tric provides the possibility to combine both intrinsic and extrinsic metrics to compare solutions that produce more reliable analyses. A set of predefined datasets, mimicking widely used three-way data and exploring crucial properties was generated and made available, highlighting G-Tric’s potential to advance triclustering state-of-the-art by easing the process of evaluating the quality of new triclustering approaches.
The paper is available: here.